|
|
@@ -120,7 +120,8 @@
|
|
|
\egroup
|
|
|
}
|
|
|
%Page de titre:
|
|
|
-\title[]{\emoji{dna} Genetic alterations in ChIP-seq}
|
|
|
+\title[]{\emoji{dna} TREC mediated oncogenesis in human immature T lymphoid malignancies
|
|
|
+preferentially involves \textit{ZFP36L2} --- Molecular Cancer}
|
|
|
|
|
|
\author{Dr. Thomas Steimlé}
|
|
|
|
|
|
@@ -154,14 +155,14 @@
|
|
|
the antigen receptor (TCR) is acquired.
|
|
|
\end{itemize}
|
|
|
\centering
|
|
|
- \includegraphics<1>[width=.4\textwidth]{Images/dev_thym.png}
|
|
|
+ \includegraphics<1>[width=.5\textwidth]{Images/dev_thym.png}
|
|
|
\includegraphics<2>[width=.6\textwidth]{Images/Tcr_str.png}
|
|
|
\end{frame}
|
|
|
|
|
|
\begin{frame}{\emoji{drop-of-blood} Background --- V(D)J recombination}
|
|
|
\begin{itemize}
|
|
|
\item<1-> This process involves a series of genome recombination of the different TCR loci
|
|
|
- followed by \alert{proliferation selection of functionnal non self-reacting receptors}.
|
|
|
+ followed by \alert{proliferation and selection of functionnal non self-reacting TCR}.
|
|
|
\end{itemize}
|
|
|
\centering
|
|
|
\includegraphics<1>[width=.7\textwidth]{Images/MaturLt.png}
|
|
|
@@ -175,7 +176,7 @@
|
|
|
\item V(D)J recombination is a \alert{threat to genomic stability},
|
|
|
prone to induce \alert{DSB occurring in genes outside of the TCR loci},
|
|
|
followed by erroneous repair resultating in SV.
|
|
|
- \item This oncogenesis process is responsible of well known genetic alterations
|
|
|
+ \item This oncogenetic process is responsible of well known genetic alterations
|
|
|
in T-ALL (particulary translocations accountable of ectopic expression of oncogenes
|
|
|
\textit{TLX1}, \textit{TAL1} etc\dots)$^1$.
|
|
|
|
|
|
@@ -206,8 +207,8 @@
|
|
|
\begin{column}{.5\textwidth}
|
|
|
\begin{itemize}
|
|
|
\item During recombination deleted parts of the loci are circulized into TRECs.
|
|
|
- \item Like tansposons the reintegration of TRECs as been suspect to cause
|
|
|
- \alert{deregulation of targeted genes}.
|
|
|
+ \item Similar to transposons, the reintegration of TRECs has been implicated in the
|
|
|
+ \alert{deregulation or inactivation of targeted genes}.
|
|
|
\end{itemize}
|
|
|
\end{column}
|
|
|
\end{columns}
|
|
|
@@ -224,14 +225,14 @@
|
|
|
\metroset{block=fill}
|
|
|
\begin{alertblock}{{\centering \large Hypothesis} }
|
|
|
\begin{itemize}
|
|
|
- \item In T-ALL, we could find with molecular biolgy tools insertions of those TRECs.
|
|
|
- \item With the same tools we also could also find all the translocations which involves the TCR.
|
|
|
+ \item In T-ALL, we could find with molecular biology tools insertions of those TRECs.
|
|
|
+ \item With the same tools we could also find all the translocations which involves the TCR.
|
|
|
\end{itemize}
|
|
|
\end{alertblock}
|
|
|
\end{frame}
|
|
|
}
|
|
|
|
|
|
-\begin{frame}{\emoji{up-arrow} Material \& Method}
|
|
|
+\begin{frame}{\emoji{triangular-ruler} Material \& Method}
|
|
|
\begin{itemize}
|
|
|
\item<1-> We used our extensed collection of \alert{T-ALL samples at diagnostic n = 1533}.
|
|
|
\item<2-> We designed a NGS capture assay with \alert{capture probes} mapped at multiple parts of de TCR \delta { } locus.
|
|
|
@@ -239,12 +240,12 @@
|
|
|
\end{itemize}
|
|
|
\begin{figure}
|
|
|
\centering
|
|
|
- \includegraphics<2>[width=.8\textwidth]{Images/Probes_TRD.png}
|
|
|
+ \includegraphics<2>[width=\textwidth]{Images/Probes_TRD.png}
|
|
|
\includegraphics<3>[width=.8\textwidth]{Images/assemblage.png}
|
|
|
\end{figure}
|
|
|
\end{frame}
|
|
|
|
|
|
-\begin{frame}{\emoji{up-arrow} Results --- \textit{TRD} translocations --- Validation cohort}
|
|
|
+\begin{frame}{\emoji{bar-chart} Results --- \textit{TRD} translocations --- Validation cohort}
|
|
|
\begin{itemize}
|
|
|
\item To validate our method, we used a previously published cohort of 264 cases analysed with \alert{\textit{TRD}
|
|
|
dual-color FISH probe}$^1$.
|
|
|
@@ -264,7 +265,7 @@
|
|
|
}
|
|
|
\end{frame}
|
|
|
|
|
|
-\begin{frame}{\emoji{up-arrow} Results --- \textit{TRD} translocations --- Discovery cohort}
|
|
|
+\begin{frame}{\emoji{bar-chart} Results --- \textit{TRD} translocations --- Discovery cohort}
|
|
|
\begin{figure}
|
|
|
\includegraphics[height=.8\textheight]{Images/distribution.png}
|
|
|
\end{figure}
|
|
|
@@ -282,7 +283,7 @@
|
|
|
}
|
|
|
\end{frame}
|
|
|
|
|
|
-\begin{frame}{\emoji{dart} Results --- \textit{TRD} translocations --- Discovery cohort}
|
|
|
+\begin{frame}{\emoji{round-pushpin} Results --- \textit{TRD} translocations --- Discovery cohort}
|
|
|
We confirmed all TRECs insertions with sanger sequencing and OGM.
|
|
|
\begin{figure}
|
|
|
\includegraphics[width=\textwidth]{Images/ZFP36L2_lolli.pdf}
|
|
|
@@ -302,8 +303,8 @@
|
|
|
any additional costs.
|
|
|
\item Our findings provide confirmation that \alert{recurrent TRECs insertions} are present in cases of
|
|
|
T-ALL.
|
|
|
- \item Using our method, we have observed that these recurrent TRECs insertions predominantly
|
|
|
- disrupt the \textit{ZFP36L2} genes.
|
|
|
+ \item Using our method, we have observed that these recurrent \alert{TRECs insertions predominantly
|
|
|
+ disrupt the \textit{ZFP36L2} gene}.
|
|
|
\item The tumor suppressor \textit{ZFP36L2} is well-established to be involved in V(D)J recombination,
|
|
|
but further clarifications are needed regarding its specific role in oncogenesis. Our findings will
|
|
|
help in the characterization of this tumor suppressor.
|
|
|
@@ -312,38 +313,152 @@
|
|
|
\end{frame}
|
|
|
}
|
|
|
|
|
|
-\begin{frame}{\emoji{dart} My PhD --- Rationnal}
|
|
|
+\begin{frame}{\emoji{thinking-face} My PhD --- Epigenetic Rationnal}
|
|
|
\begin{itemize}
|
|
|
- \item The aforementioned results contribute to the understanding of the dysregulation of proto-oncogenes
|
|
|
+ \item<1-> The aforementioned results contribute to the \alert{understanding of the dysregulation of proto-oncogenes}
|
|
|
such as \textit{TAL1}, \textit{TLX1} or \textit{TLX3}.
|
|
|
- \item Despite extensive investigation, the molecular mechanisms underlying the dysregulation of these oncogenes,
|
|
|
- remain elusive in many cases.
|
|
|
- \item
|
|
|
+ \item<2-> Despite extensive investigation, \alert{the molecular mechanisms} underlying the dysregulation of these oncogenes,
|
|
|
+ \alert{remain elusive in many cases}.
|
|
|
+ \item<3-> It has been demonstrated that \alert{tumor cells acquire enhancers}\footnotemark[1] through intergenic
|
|
|
+ sequence mutations that enable binding of transcription factors.
|
|
|
+ \end{itemize}
|
|
|
+ \begin{figure}
|
|
|
+ \centering
|
|
|
+ \includegraphics<3>[width=.75\textwidth]{Images/bradner_cis_small.png}
|
|
|
+ \end{figure}
|
|
|
+ \footnotetext[1]{Bradner JE, et al. Cancer. Cell. 2017 Feb 9;168(4):629-643}
|
|
|
+\end{frame}
|
|
|
+
|
|
|
+\begin{frame}{\emoji{thinking-face} My PhD --- Epigenetic Rationnal}
|
|
|
+ \begin{itemize}
|
|
|
+ \item The presence of upstream \alert{indels in \textit{TAL1} leads to the formation of a neo-enhancer}$^1$.
|
|
|
+ \item It has also been shown that the transcription factor \alert{MYB can bind to this neo-enhancers}$^2$.
|
|
|
+ \end{itemize}
|
|
|
+ \begin{figure}
|
|
|
+ \includegraphics[width=.46\textwidth]{Images/tal_ins.png}
|
|
|
+ \end{figure}
|
|
|
+ \footnotetext[1]{
|
|
|
+ \tiny
|
|
|
+ Navarro JM et al. Nat Commun. 2015;6:6094}
|
|
|
+ \footnotetext[2]{
|
|
|
+ \tiny
|
|
|
+ Smith, C et al. “TAL1 activation in T-cell acute lymphoblastic leukemia: a
|
|
|
+ novel oncogenic 3' neo-enhancer.” Haematologica vol. 108,5 1259-1271. 1 May. 2023}
|
|
|
+\end{frame}
|
|
|
+
|
|
|
+{\setbeamercolor{background canvas}{bg=bgturq}
|
|
|
+\begin{frame}[c]
|
|
|
+ \vspace{.6cm}
|
|
|
+ \metroset{block=fill}
|
|
|
+ \begin{alertblock}{{\centering \large Hypothesis} }
|
|
|
+ \begin{itemize}
|
|
|
+ \item[A]<1-> By taking a pan-genomic approach, it is possible to identify numerous \alert{intergenic
|
|
|
+ alterations correlated with the cis deregulation of adjacent genes} (correlation between
|
|
|
+ transcriptome RNA-seq and ChIP-seq).
|
|
|
+ \item[B]<2-> Some of these genes are expected to be known oncogenes, while others have
|
|
|
+ the potential to be \alert{novel oncogenes} (discovery).
|
|
|
+ \item[C]<3-> It is likely that these specific alterations are the causative factors
|
|
|
+ for the cis deregulation of adjacent genes (functional experiments).
|
|
|
+ \item[D]<4-> Based on these alterations, it is possible to stratify patients into
|
|
|
+ groups with similar \alert{prognoses}.
|
|
|
+ \item[E]<5-> The \alert{characterization of the deregulation mechanisms and the discovered oncogenes}
|
|
|
+ should help identify vulnerabilities that can be targeted by treatment.
|
|
|
+ \item[F]<6-> This treatment may prove to be \alert{more effective with fewer side effects} compared
|
|
|
+ to the currently prescribed polychemotherapy.
|
|
|
+ \end{itemize}
|
|
|
+ \end{alertblock}
|
|
|
+\end{frame}
|
|
|
+}
|
|
|
+
|
|
|
+\begin{frame}{\emoji{white-check-mark} What's already done}
|
|
|
+ \begin{itemize}
|
|
|
+ \item[\emoji{white-check-mark}] Alignement and copy number analysis of more than 260 RNA-seq
|
|
|
+ \end{itemize}
|
|
|
+ \begin{figure}
|
|
|
+ \includegraphics[width=\textwidth]{Images/fdt_hm.png}
|
|
|
+ \end{figure}
|
|
|
+\end{frame}
|
|
|
+
|
|
|
+\begin{frame}{\emoji{white-check-mark} What's already done}
|
|
|
+ \begin{itemize}
|
|
|
+ \item[\emoji{white-check-mark}]<1-> Alignement and copy number analysis of more than 260 RNA-seq
|
|
|
+ \item[\emoji{white-check-mark}]<1-> Alignement and copy number analysis of 72 ChIP-seq (H3K4me4 and H4K27ac).
|
|
|
+ \item[\emoji{white-check-mark}]<2-> We have developed a high-performance tool that optimizes memory usage and
|
|
|
+ speed for correlating variably expressed genes with depth of ChIP-seq peaks. Specifically, our tool
|
|
|
+ efficiently handles a large dataset consisting of 10,036 gene expressions and 84,839 ChIP-seq positions,
|
|
|
+ resulting in a total of 851,444,204 correlations.
|
|
|
+ \item[\emoji{white-check-mark}]<3-> Calling of genetic alterations sequenced by ChIP-seq.
|
|
|
+ \end{itemize}
|
|
|
+\end{frame}
|
|
|
+
|
|
|
+\begin{frame}{\emoji{dart} First results}
|
|
|
+ With these filters:
|
|
|
+ \begin{itemize}
|
|
|
+ \item Recurrent mutations in the same ChIP-seq peak (> 1 case) with enrichment of the alternative allele (AF > 0.6).
|
|
|
+ \item Cases with the mutations should have correlated genes (Pearson coefficient > 0.7)
|
|
|
+ significantly upregulated (t-test p value < 0.05)
|
|
|
+ \end{itemize}
|
|
|
+ \begin{figure}
|
|
|
+ \includegraphics[width=\textwidth]{Images/hm_1.png}
|
|
|
+ \end{figure}
|
|
|
+ \vspace{.5cm}
|
|
|
+ \begin{figure}
|
|
|
+ \includegraphics[width=\textwidth]{Images/hm_2.png}
|
|
|
+ \end{figure}
|
|
|
+\end{frame}
|
|
|
+
|
|
|
+\begin{frame}{\emoji{knocked-out-face} Caveats}
|
|
|
+ \begin{itemize}
|
|
|
+ \item<1-> Most of intergenic alterations are \alert{SNPs}.
|
|
|
+ \item<2-> Complexe alterations like \alert{indels and SV are difficult to call with ChIP-seq small reads}.
|
|
|
+ \item[$\rightarrow$]<3-> We are implementing \alert{longreads sequencing} with the Oxford Nanopore Promethion for
|
|
|
+ resolving complex genomic regions, detecting structural variations, and studying repetitive elements.
|
|
|
+ \end{itemize}
|
|
|
+ \begin{figure}
|
|
|
+ \includegraphics<3>[width=.4\textwidth]{Images/promethion.png}
|
|
|
+ \end{figure}
|
|
|
+\end{frame}
|
|
|
+
|
|
|
+\begin{frame}{\emoji{flying-saucer} Longreads pipeline}
|
|
|
+ \begin{itemize}
|
|
|
+ \item<1-> We will conduct a \alert{whole-genome sequencing of 150 T-ALL cases along with their corresponding constitutional samples}.
|
|
|
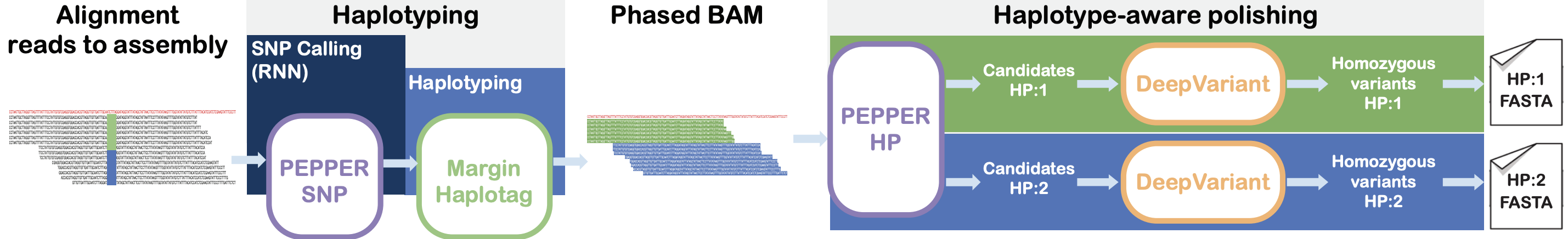
+ \item<2-> Our objective is to implement a pipeline capable of \alert{assembling long reads and generating a diploid
|
|
|
+ reference genome} for each of the cases.
|
|
|
\end{itemize}
|
|
|
+ \begin{figure}
|
|
|
+ \includegraphics<2->[width=\textwidth]{Images/lr_pipe.png}
|
|
|
+ \end{figure}
|
|
|
+
|
|
|
+ \footnotetext[1]{
|
|
|
+ \tiny
|
|
|
+ Kolmogorov, Mikhail et al. “Assembly of long, error-prone reads using repeat graphs.” Nature biotechnology
|
|
|
+ vol. 37,5 (2019): 540-546. doi:10.1038/s41587-019-0072-8}
|
|
|
\end{frame}
|
|
|
|
|
|
-\begin{frame}{\emoji{dart} Troisième année de thèse -- Hypothèses E et F}
|
|
|
+\begin{frame}{\emoji{flying-saucer} Longreads pipeline}
|
|
|
\begin{itemize}
|
|
|
- \item Selon les oncogènes identifiés lors des étapes précédentes nous explorerons les possibles interventions thérapeutiques
|
|
|
- \item Possibilité de test phramacologiques \textit{ex vivo} et \textit{in vivo}.
|
|
|
+ \item<1-> We will conduct a \alert{whole-genome sequencing of 150 T-ALL cases along with their corresponding constitutional samples}.
|
|
|
+ \item<1-> Our objective is to implement a pipeline capable of \alert{assembling long reads and generating a diploid
|
|
|
+ reference genome} for each of the cases.
|
|
|
+ \item<1-> The alignment of somatic reads on it and the subsequent calling of somatic variants,
|
|
|
+ especially \alert{structural variations} (SV), will be of significant interest.
|
|
|
+ \item<2-> By aligning our current RNA-seq and ChIP-seq data using this approach, we will be able to \alert{phase gene
|
|
|
+ expression} and identify \alert{allele-enriched epigenetic marks} more efficiently.
|
|
|
+ \item<3-> We will also have access to \alert{phased methylation of CpG islands} with the same technic.
|
|
|
\end{itemize}
|
|
|
\end{frame}
|
|
|
|
|
|
% qrencode https://git.t0m4.fr/Thomas/presentation_projet_inserm/raw/master/presentation.pdf -t SVG | sed 's/"#000000"/"#e3b505"/g' | sed 's/"#ffffff"/"#23373B"/g' | /Applications/Inkscape.app/Contents/MacOS/inkscape -p Images/qr-code.pdf
|
|
|
\begin{frame}[standout]
|
|
|
- \alert{\textbf{Questions ?}}\\
|
|
|
- \vspace*{1.5cm}
|
|
|
- \emoji{calling}
|
|
|
+ \vspace*{1cm}
|
|
|
+ \textcolor[HTML]{e3b505}{Thank you for listening !}\\
|
|
|
+ \textcolor[HTML]{e3b505}{\textbf{Questions ?}}\\
|
|
|
+ \vspace*{1cm}
|
|
|
\begin{figure}
|
|
|
\includegraphics[width=.2\textwidth]{Images/qr-code.pdf}
|
|
|
\end{figure}
|
|
|
\end{frame}
|
|
|
|
|
|
-% \begin{frame}
|
|
|
-% \frametitle{\emoji{floppy-disk} Méthodes > Calling > Lancet}
|
|
|
-% Source code : \url{https://github.com/nygenome/lancet}
|
|
|
-% \vskip 0.2in
|
|
|
-% \lstinputlisting[language=bash, caption={lancet -- bash version}, style=mystyle]{Codes/lancet.txt}
|
|
|
-% \end{frame}
|
|
|
+% qrencode https://git.t0m4.fr/Thomas/presentation_projet_inserm/raw/master/presentation.pdf -t SVG | sed 's/"#000000"/"#e3b505"/g' | sed 's/"#ffffff"/"#23373B"/g' | /Applications/Inkscape.app/Contents/MacOS/inkscape -p --without-gui --export-pdf=Images/qr-code.pdf
|
|
|
|
|
|
\end{document}
|